 Bbedit clippings code#This code can be applied to any molecule in a sphere representation. Grayscale is not an option available through a pulldown in PyMOL. This variant of ambient occulusion colors the carbon atoms black and then applies grayscale. This code can be applied to any molecule in the sphere representation. 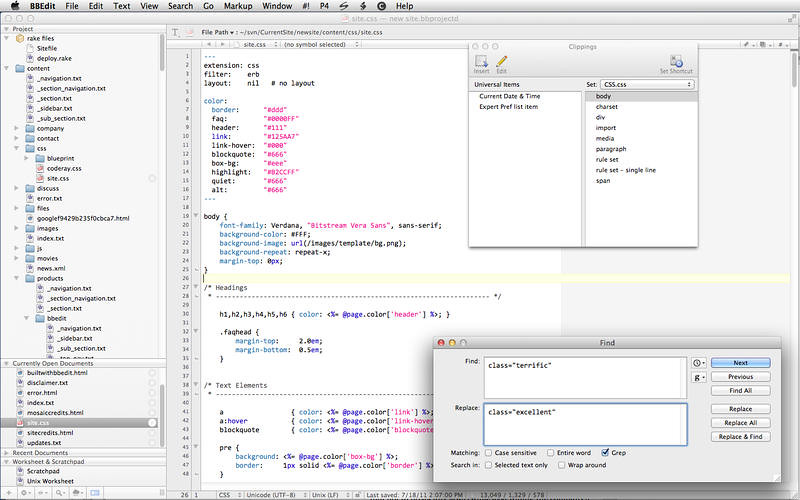
This is the grayscale verison of ambient occlusions. This variant of ambient occulusion colors the carbon atoms black. Enter hide spheres to remove the spheres. It applies ray tracing so moving the molecule in the viewport ruins the effect. This code can be applied to any molecule in the sphere representaton. The effect is not available via a GUI pulldown mean. You may have to refresh the web browser several times to get all of the images to display.Īpplies a photorealistic effect. Here are of some figures that are impossible or tedious to make via the PyMOL GUI alone. (Use the spse function in the pymolshortcuts.py file in the pymolshortcuts repository to save session files with time stamps to avoid overwriting previously saved session files.) pml file extension, to an opened log file, or to a frequently saved session file, the work can be lost. If the commands are not saved to an open script file with a. It is difficult to issue so many commands through PyMOL's graphical user interface (GUI) without making mistakes. Over 100 lines of pml commands and settings are required to make more sophisticated images. The macro language sends arguments to Python functions, but its syntax is simpler for non-programmers to understand than the syntax of Python code. These scenes can be made into static images for posters, seminars, and manuscripts, or they can serve as parts of molecular movies. The PyMOL macro language (pml) can be used to set parameter values and execute commands to make customized scenes of biomolecules in PyMOL's viewport. The PyMOL GUI is useful for making the images of global scenes, but PyMOL rapidly becomes tedious to use to make images of detailed scenes. PyMOL is also popular for making movies of molecules. PyMOL is often used to make cover images for scientific journals. PyMOL's vast array of parameters provides exquisite control over the appearance of the output. PyMOL is a leading molecular graphics program for making images of proteins and nucleic acids for publication. 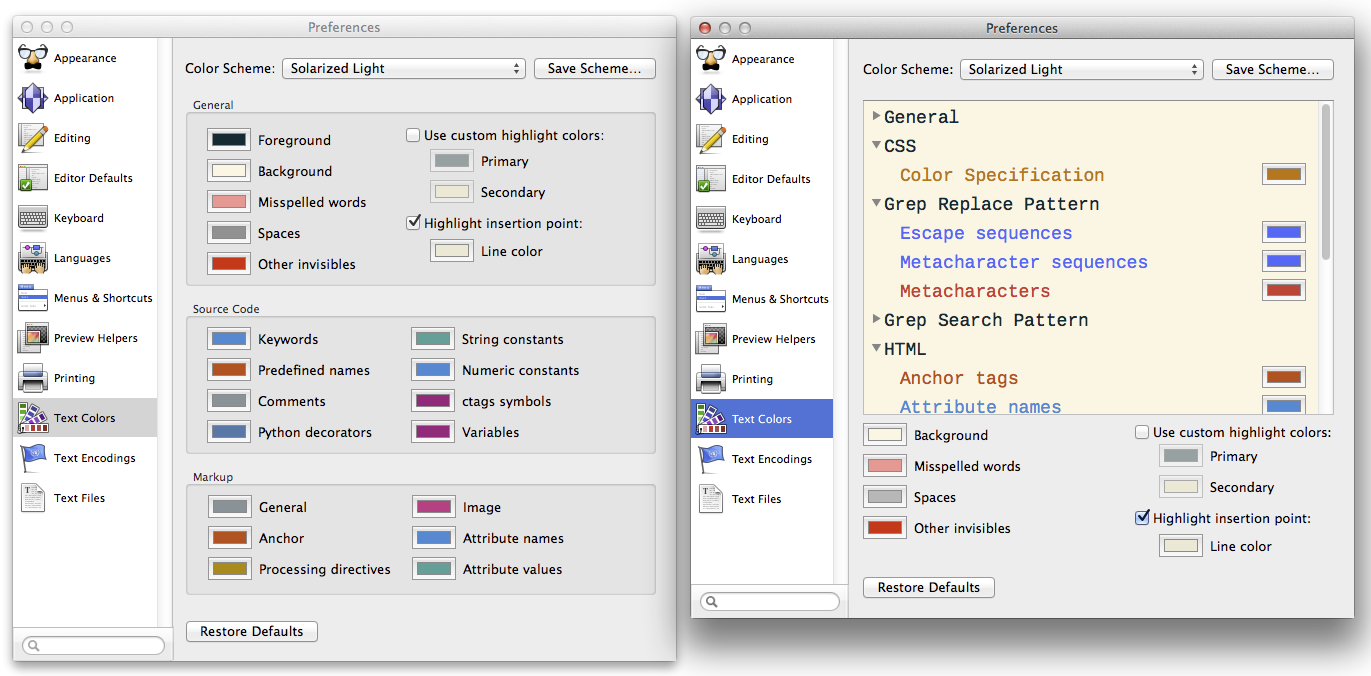
Go back to the MooersLab/pymolsnips GitHub page
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
